Week 5: ASReview and More Screening
April 13, 2026
Hello everyone! For this week’s updates, I’ll again mainly be focusing on my independent research outside of my internship, since Dr. Iyer has been out this week and I’ve been taking a break as a result. No worries, though, since I’ll be returning next week and bringing more updates. As a follow-up from last week, I reached out to my father and finally got ASReview to work on my computer, but, as a result, I’ve had to redo screening (again) with harsher parameters. I started screening really early, though, so as a bonus, I still have enough leeway to handle small setbacks like these. Screening again did take me around 2 workdays, so I didn’t get to the data extraction (as I had hoped I’d get to), but using an AI-assisted data extraction program on R (an open-source programming language and environment specializing in statistical computing, data analysis, and graphics) should streamline the process.
Of course, as with using any AI tools, I’ll be double/triple-checking everything to ensure the data is correct, but it serves to help make the extraction marginally faster. This whole process has been just learning as I go on, so mistakes are bound to happen, but I am doing my best at logging everything and tracking all the changes I make so my study will fit meta-analysis standards. With ASReview, I still screen everything by hand, but it is much easier to separate texts since they have a really intricate tagging system, which allows me to annotate for studies I want to go back and reference mine or studies that I can’t use for my analysis, but would still be good to include in my literature review. It also keeps track of the intervals between each study marked “relevant” and records data during the screening process (all of which I’ve attached below).

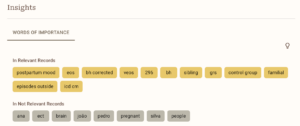
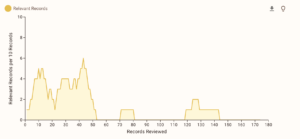
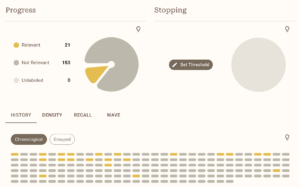
I do have a small sample of studies (mainly because my topic is not heavily explored yet), so I do have some concerns about that, but to combat that, I am trying to gain access to more databases and go through the references of some other meta-analyses and systematic reviews. I have high hopes going into data extraction and am looking to have at least one model in the works for next week to share. Below is a summary of all the exclusion criteria and inclusion criteria I used for (hopefully) my last time screening.
Inclusion Criteria:
In order for studies to be included in this analysis, they must:
- Must contain enough information to calculate an effect size
- Must be published in English
- Must be publicly available
- Must examine the relationship between Schizophrenia and postpartum disorders
Exclusion Criteria:
Studies were excluded from the analysis if:
- Did not fit the inclusion criteria
- They do not include a clinical or validated diagnostic measure of either postpartum illness or schizophrenia spectrum disorder (e.g., studies relying solely on self-report without clinical confirmation, or that use non-standardized definitions of postpartum illness such as general “postnatal distress”)
- The focus was on the effects of a medication used to treat both disorders (or neither of the disorders at all—i.e., wrong focus)
- The population used in the study was not human (i.e, mice, etc)
- Duplicate/same dataset as another included study
- Study type: Case report, editorial, qualitative only, review/meta-analysis
Till next time,
Yalini Kathamuthu

Leave a Reply
You must be logged in to post a comment.