Week 6 (3/30-4/3) - Testing Baseline ML Models for ICD Classification (And Why They Failed)
April 3, 2026
Welcome back to my blog! This week was a very humbling one. As I finished training the baseline models, the results had a clear and unfortunate conclusion: models like Logistic Regression, Random Forest, and Bayesian Logistic Regression are simply not built for the complexity of this problem.
My first priority for this week was tackling the logistic regression performance, seeing how much I could fix the overfitting. I tried three different approaches. First, I dropped all rows that didn’t have any of the top 59 ICD titles, which reduced the dataset from 126 to 106 rows. Training accuracy was 95%, but test accuracy dropped even further to about 4.5%. Next, I reduced the label space even more aggressively to only the top 16 titles (those appearing at least 4 times), leaving just 65 rows. Training accuracy remained high at 96%, but test accuracy fell to 0%. These approaches didn’t improve test performance, likely due to the large amount of rows not included with each filtering.
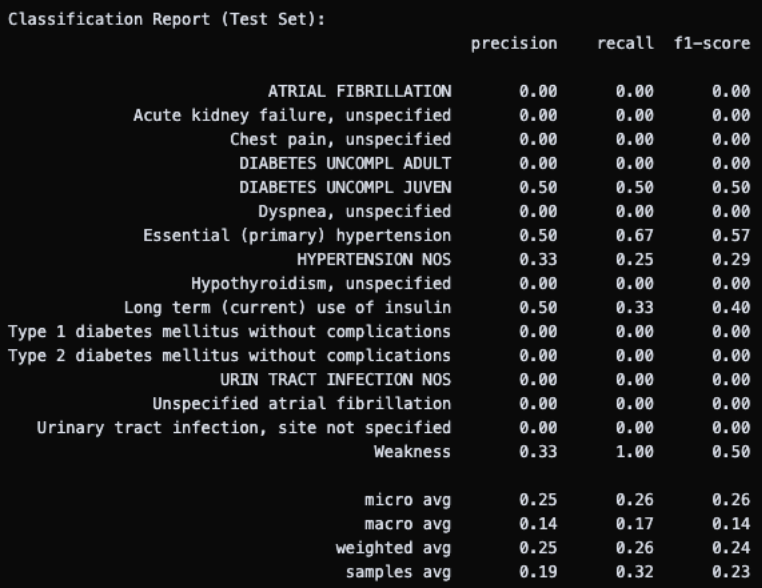
Finally, I tried grouping ICDs into their broader categories to reduce the label space while preserving some clinical meaning (though this was not ideal). Training accuracy dropped to a more reasonable 76%, but test accuracy was still completely zero. At this point, it was clear that reducing the label space wasn’t fixing the fundamental problem, convincing me further that the model was just structurally too simple for this kind of high-dimensional, sparse multilabel problem.
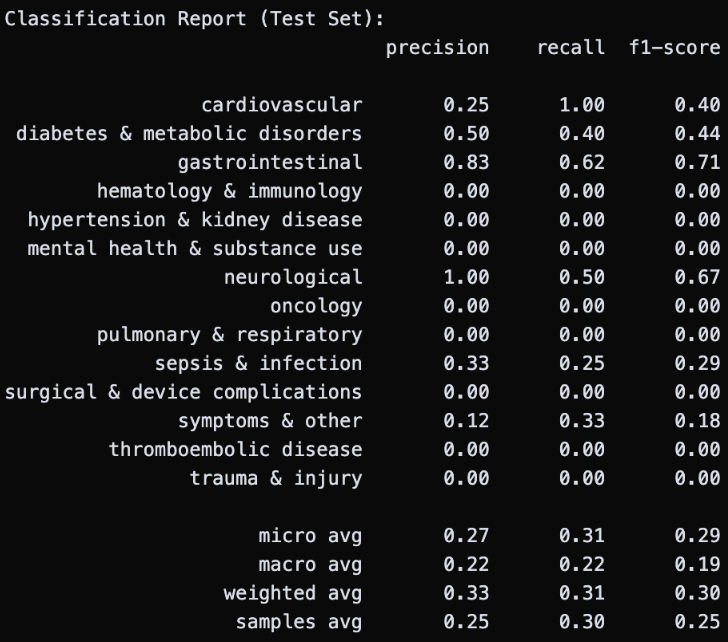
Then, I tried to implement a Random Forest using the same MultiLabelBinarizer and MultiOutputClassifier setup, to see if it did any better. Unfortunately, the performance was again disappointing, with massive overfitting and zero test accuracy (1.0 train accuracy!). To fight this, I constrained the trees directly: setting a maximum depth of 5, requiring at least 5 samples per leaf, and at least 10 samples before each split. This brought training accuracy down to 0.83, a promising sign that the model was no longer just memorizing the training data, but this was not enough as test accuracy remained at zero.
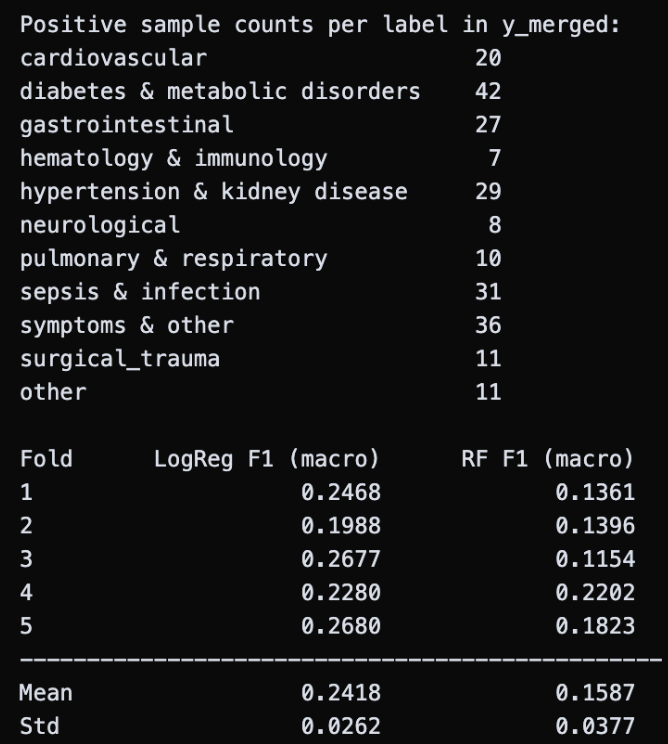
That’s when I had the idea to apply cross-validation to both models. Rather than relying on a single train-test split, cross-validation rotates through multiple subsets of the data (5-fold meaning five different splits), giving a much more reliable picture of true performance without overfitting. I also merged a few smaller ICD categories together, consolidating the three categories with fewer than 6 total occurrences and combining surgical & device complications with trauma & injury to reduce noise. The results were definitely better than the zeros I had been seeing before: logistic regression achieved a mean F1 score of 0.24, and random forest reached 0.16. However, they are still not clinically meaningful, reinforcing my original hypothesis. Both models are too simple to capture the complex, interdependent relationships between features that clinical reasoning requires. While disappointing, this takeaway actually showcases the need for the project’s more powerful approach of an LLM.
With the classical baselines exhausted, I started exploring Bayesian logistic regression. As I covered in my reading a few weeks ago, Bayesian updating lets you combine a prior belief with observed evidence to produce a posterior distribution over predictions, rather than a single point estimate. This is particularly well-suited to this problem: it handles class imbalance more gracefully, is more robust to overfitting with limited data, and provides interpretable credible intervals. While I didn’t hold a lot of hope, I wanted to confirm the performance before moving on to training the LLM. For my model, I assumed a prior of 0 for each feature coefficient as built-in regularization, and modeled each ICD label as a Bernoulli outcome since each is either applied or not.
My first attempt was to use the PYMC library, the standard tool for probabilistic programming in Python, but I kept running into installation issues even after resolving several dependency conflicts between libraries. So, as a workaround, I implemented a scikit-learn approximation: fitting logistic regression with L2 regularization across many bootstrap-resampled versions of the training data. In the end, it provides an excellent approximation of the results (posterior distribution of coefficient values) of an ideal Bayesian model from the PYMC library. Unfortunately, the runtime for this (with cross-validation) is much longer and so I was not able to include the results in this blog post, but I’ll dive into them next week!
In parallel, I’ve been working on installing GPT-OSS, the open-source language model I plan to fine-tune. I ran into a major compatibility issue with the torch library required to load models from HuggingFace. Investigating and debugging the problem, I found that torch couldn’t be installed with the Python version I had been using, requiring me to downgrade from Python 3.13 to 3.12. This allowed me to successfully install the model, which I’m very excited to start training next week!
If you’ve worked with Bayesian approaches on small or imbalanced datasets, I’d love to hear your experience and whether you faced similar issues. For next week, I plan on training GPT-OSS with the CREST corpus and the MIMIC-IV dataset, finally reincorporating the information that was lost in featurizing for the baseline models and (hopefully!) getting much clearer and clinically accurate results. Stay tuned for the blog!
Reader Interactions
Comments
Leave a Reply
You must be logged in to post a comment.

Hi Aanya, it sucks when the model training process slows everything down more than expected. That’s where I was 2 weeks ago. And the fact that no matter what you do, it’s still a bit of a black box can be discouraging. It’s good that you still verified yourself that the simpler models won’t work, rather than just assuming. I think you’re on the right track by trying more complex approaches. Looking forward to the results!
Hi Anav, thank you – it’s reassuring to hear that you’ve had similar experiences (I think anyone who’s worked with machine learning models before can relate)! Looking forward to hearing your thoughts on this week’s results.
Hi Aanya, great work this week! Even though the original models you tried didn’t end up working out, I think that trying them out was still a valuable part of the process. Hope to see more progress next week!
Hi Elin, thank you so much! Yes, I am very excited for this week’s work – looking forward to seeing your thoughts on it!
Hi Aanya, good progress. It sucks that tuning wasn’t the only issue. I like the idea of moving from baseline models to cross validation and then bayesian approaches. I am excited to see how GPT-OSS works for you.
Hi Aariv, thank you! Yes, I’m glad I tried the different baseline models and cross validation first (even before hyperparameter tuning) so I knew I had exhausted a breadth of options and taking the more complex LLM approach was the right step. Looking forward to hearing your thoughts on this week’s work!