Week 2: Literature Review & PyHFO Deep Dive
March 20, 2026
Welcome back to the blog, everyone! This week, I’m diving into my literature review and showing how to use PyHFO, and what I encountered while researching and identifying specific SYNGAP1 biomarkers. I also really enjoyed answering all of your thoughtful questions from last week.
This week’s literature review centered on two key papers: a 15-page paper on PyHFO 2.0: An Open-Source Platform for Deep Learning–Based Clinical High-Frequency Oscillations Analysis, and a 120-page foundational reference called EEG for Beginners, which covers everything from basic signal interpretation to complex epilepsy analysis.
In the initial paper surrounding the deep learning pipeline and framework of PyHFO, it introduces PyHFO as a tool that integrates classical signal processing with modern deep learning for automated HFO (High-Frequency Oscillation) biomarker analysis. Since PyHFO requires a graphical interface, it must be run through a virtual desktop environment on a server, and the paper walks through how to set this up for shared use. The deep learning models are sourced from the Hugging Face ecosystem, a platform hosting thousands of pre-trained models, and are specifically configured for two classification tasks: spkHFO detection for identifying spike-associated HFOs, and eHFO identification for identifying epileptogenic HFOs. Importantly, this implementation of PyHFO is dedicated to analyzing EEG recordings from mice with epilepsy, which is directly relevant to my SYNGAP1 research.
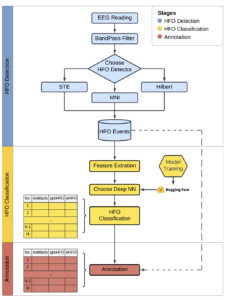
In the image above, the paper below illustrates the full PyHFO pipeline. EEG data enters the system and passes through a BandPass Filter, after which the user selects an HFO detector, either STE, Hilbert, or MNI. Detected HFO events are then passed to a Feature Extraction step, followed by a Deep Neural Network classifier loaded from Hugging Face that performs HFO Classification into spkHFO and eHFO categories. The final stage is Annotation, where results are stored and reviewed.
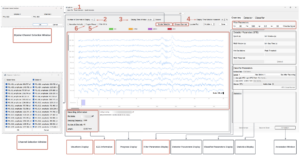
From there, above is the graphical interface of pyHFO and how it is possible to run detections and classifications of mice EEG data, and now let me show how I apply it to the mice datasets. In the top left, I can upload an EDF file (the standard file format for EEG recordings) from our SYNGAP mouse directory and then select specific filter parameters. I then can run a Quick Detection pass over the EEG scan to get a baseline read of HFO events in the recording. From there, in the right panel, I can configure specific detection parameters and re-run detection, which will return a precise count of HFOs detected within the EDF file. After detection, I will press HFO Classification, which outputs a statistical breakdown of spkHFOs, eHFOs, and total HFOs across the recording in the bottom right panel. Finally, I can annotate the detected HFOs for more analysis.
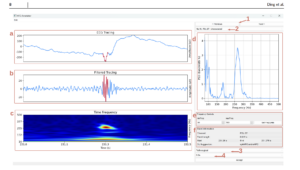
Then from there, there is an annotation window provides four synchronized views for each detected HFO event. Panel (a) shows the EEG Tracing, which is the raw unfiltered signal. Panel (b) shows the Filtered Tracing, which is the bandpass-filtered signal highlighting the HFO. Panel (c) shows the Time-Frequency Plot, a spectrogram showing the HFO’s energy distribution across time and frequency. Panel (d) shows the FFT Plot, the Fast Fourier Transform showing the dominant frequency components of the event. Together, these views allow me to precisely classify spike morphology and verify whether a detected event is a true spkHFO or eHFO rather than an artifact.
And now, let me dive into the second paper of our initial literature review for the EEG for Beginners. It teaches the essential groundwork for interpreting EEG signals across different brain states. For example, when awake with eyes closed, the dominant rhythm is posterior alpha (8-12 Hz), maximal at electrodes O1/O2, and it immediately attenuates when the eyes open, which is a key marker of a healthy awake brain. Low-voltage beta activity is also visible anteriorly. In the drowsy state, alpha disappears and is replaced by theta activity, and normal drowsy features include vertex sharp waves at Cz and positive occipital sharp transients of sleep (POSTS), both of which are normal and not epileptiform, though beginners often flag them as abnormal. Then in light sleep (Stage 2), two highly specific normal patterns appear: sleep spindles (12-14 Hz bursts lasting 0.5-3 seconds at central electrodes) and K-complexes (high-amplitude slow waves often preceded by a vertex sharp), both of which are pathognomonic for Stage 2 NREM sleep. In deep sleep (Stages 3-4), high-amplitude delta activity comprising at least 20% of the epoch dominates, and the EEG appears chaotic and slow. Across all of these, it shows images of EEG scans of these scenarios that allow to track patterns in order to analyze and interpret EEG signals in these different brain states.
Then, across from that, beyond normal brain states, the paper also covers how to identify seizure patterns from spikes. The image below illustrates four major seizure morphologies: rhythmical spiking, rhythmically evolving theta/delta activity, spike-wave complexes, and electrodecremental patterns (low voltage fast). The accompanying EEG sample to the right also shows an example of how it teaches to analyze many types of spikes in EEG scans as it shows a left superior frontal seizure beginning with rhythmical spiking.

So this week’s blog covered everything I learned through my literature review, from understanding and analyzing EEG scans to a deep dive into PyHFO and its future application to our SYNGAP1 mice datasets. For next week, I will be showing a real walkthrough of debugging an issue in pyHFO followed by a complete detection and classification run across all EEG files in the SYNGAP mouse directory. It has been genuinely exciting to learn how to apply and fine-tune PyHFO to extract meaningful biomarker data from these datasets. Alongside for next week, I’ll also be sharing how I plan to integrate Claude Code to accelerate my ML model runs and streamline debugging going forward!
Reader Interactions
Comments
Leave a Reply
You must be logged in to post a comment.

Hi Dhruv! I like how you connected each paper to your project really clearly and went through each paper clearly with so much detail. One question I had was are there any limitations of PyHFO that stood out to you, and have you predicted any obstacles and/or solutions to getting over them? I’m especially interested in seeing how you implement Claude Code in your project, as I’ve noticed many of our fellow Senior Project peers are using AI to help them code faster and more effectively!