Week 3: Debugging the PyHFO Codebase
March 31, 2026
Welcome back again to my blog everyone! This week I’m diving into an issue that I ran into while trying to run mice EEG datasets through pyHFO, and how I was able to use coding agents such as Claude Code to assist me in debugging the issues and fix the root causes.
As I explained last week, pyHFO is a tool for detecting and classifying these High-Frequency Oscillations (HFOs) in EEG data. HFOs are brief bursts of electrical activity in the brain that researchers use as biomarkers, biological signals that can indicate neurological conditions. So when I tried loading a mice dataset, the tool would find HFOs just fine (23 HFOs in my image of test file below), but then freeze when trying to classify them. With hundreds of mice datasets to process, a tool that freezes halfway through was a major problem that needed to be fixed. The pyHFO classification models were “hanging”, essentially meaning the classifier was never completing, so no HFOs were being sorted into their correct categories and were left everything unlabeled.
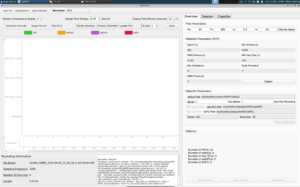 In the image above, it shows the problem. When we try to open mice file and load a quick detection of a mice dataset, the tool is only able to detect the total number of HFOs, as seen in the bottom right showing 23 HFOs, but unfortunately is unable to classify which ones are artifacts, real HFOs, spkHFOs, or eHFOs. As seen on the left, it also isn’t able to display the EEG waves in the window, leaving it blank and frozen with nothing to annotate. And this is just one mice dataset, with hundreds more that need to be classified, showing that this hang needed to be fixed immediately.
In the image above, it shows the problem. When we try to open mice file and load a quick detection of a mice dataset, the tool is only able to detect the total number of HFOs, as seen in the bottom right showing 23 HFOs, but unfortunately is unable to classify which ones are artifacts, real HFOs, spkHFOs, or eHFOs. As seen on the left, it also isn’t able to display the EEG waves in the window, leaving it blank and frozen with nothing to annotate. And this is just one mice dataset, with hundreds more that need to be classified, showing that this hang needed to be fixed immediately.
So now, lets start off with how I fixed this, first I was able to incorporate the coding agent Claude Code that gave me ideas of where there could be bugs in the codebase that may need to be fix. Using Claude Code to navigate the codebase, I found that when classification started, the program was trying to download its AI models from the internet (a platform called HuggingFace, which hosts machine learning models that I explained in our previous blog about how pyHFO works) before checking if copies were already saved locally on the machine. With no internet access as it is on a server, it would just wait forever for a download that was never coming with no timeout and no error message. The fix was to tell the program to always check for local model files first, and only go online if nothing is found locally.
After figuring that out and fixing that, a another new error appeared saying the program couldn’t find a module called “src”. Essentially, the problem was that the locally saved model files were created back when the software package was named “src”. The package was later renamed to “pyhfo2app”, but the saved files still had the old name baked into them. So when the program tried loading in those files, it went looking for “src”, and then couldn’t find it, and crashed. The fix was to add a small redirect in the code that says “if you’re looking for “src”, it’s actually called “pyhfo2app” now, essentially this alias so the old files could still be read correctly.
Then the next error came: “Please load artifact model first!”, even though the models had just been loaded. The real issue was that the program had these three internal variables (model_a, model_s, model_e) that were never given a starting value of “empty” when the program launched. So when the code checked whether a model was loaded by asking “is this variable empty?”, it crashed because the variable didn’t exist at all yet, like asking for the value of a box that was never put on the shelf. The fix was simply to initialize all three variables to None (meaning “nothing loaded yet”) right at startup when running a quick detection for pyHFO. From there, during its GUI testing (Graphical User Interface), classification would reach 0% and freeze again. This time though with a warning about timers being started from the wrong thread. To understand this, the program was trying to practically speed things up by running multiple tasks simultaneously with parallel processing. However, once a graphical interface like the pyHFO window or a GPU has been started up, bringing new parallel processes causes a deadlock, and nothing moves forward. So the fix was to detect when this situation could occur and switch to running tasks one at a time instead of in parallel, sacrificing a little speed to avoid the freeze entirely.
After finding and fixing all four of these errors, everything eventually worked. I loaded the test file, ran detection (23 HFOs found), and classification completed successfully, sorting them into 19 artifacts, 4 real HFOs, 1 spkHFO (an HFO occurring alongside a spike in brain activity), and 3 eHFOs (a specific subtype of HFO). The GUI displayed “Classification finished!” and the EEG waveforms to the left rendered fully and were ready to annotate, as shown in the image below.
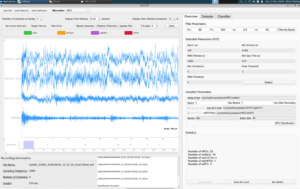
So this week’s blog covered the process in how I used Claude Code as this debugging agent to systematically work through a tricky, multi-layered bug in the pyHFO codebase. For next week, I will be diving deeper into now running the detector and classifier, and maybe even analyzing the EEG waveforms across a full batch of SYNGAP1 files, getting closer to meaningful results in biomarker development in this research!
Reader Interactions
Comments
Leave a Reply
You must be logged in to post a comment.

Hi Dhruv, amazing progress this week! I really liked how patiently you went through each error (which could be easily frustrating!), brainstorming and addressing the cause. I also liked how you used Claude Code to do this efficiently while ensuring you still understood the root of each problem. Do you think you’ll continue to implement Claude Code as your project progresses, and at the same scale?