Week 5: Fixing Bugs on the PyHFO Codebase
April 15, 2026
Hey everyone, welcome back to my blog! This week, I was able to successfully run the HFO detector on the Syngap files and save the output data in the same folder as the EDF files (so that we can load them back in and run classification once we troubleshoot that step), fix a processing issue in the PyHFO codebase that was slowing down the analysis of longer mouse recordings, and continue my literature review by reading a research paper on a nine-day continuous EEG recording study alongside two hours of high-density EEG under chronic sleep restriction conditions in mice.
Starting off, the first main task this week was running the STE (Short-Term Energy) detections manually across all the Syngap files and saving the output data in the same folder where the EDF files live, without including the NPZ files, since those act like duplicates and clutter things up. EDF and NPZ, by the way, are some of the standard file formats used for storing EEG recordings, and keeping all the outputs organized alongside them is important for the next classification step. From there, in the first image below, you can see the set of Syngap post-treatment EDF files at postnatal day 120 (P120) being stored and organized in the codebase. The bottom section of the terminal shows all the post-treatment EDF files that are queued up and going to be used in the detections, and you can also see the cleanup process happening in real time, where we’re working to remove those NPZ duplicate files to keep the directory clean and ready to go for detections.
After that, however an huge issue was coming up with the file size. The long recordings, anywhere from four to six hours, were causing the pipeline to hang and fail to process correctly, not letting the detections run. On all the shorter ten-minute files, the detections ran completely smoothly without a hitch, but on the longer segments it would slow way down and eventually stall out. So the solution was clear: we needed to slice those longer EDF files into shorter segments before running the detector on them.
To solve this, we wrote a quick shell script to do exactly that: “for ff in data/*.edf; do python scripts/edf_segment.py –start 0 –duration 600 -o $ff.10min.edf $ff; done”. This loops through every EDF file in the data folder, calls a Python segmentation script, and outputs a clean ten-minute (600-second) version of each file, and it worked. The hang was completely gone through making shorter file for a short-term solution of running the longer recordings.
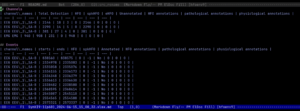
In the second and third images above, you can see the results of those shorter ten-minute detections running smoothly on the Syngap p60 files. The output shows channel-level statistics with total detections, HFO counts, spkHFO, eHFO, and event-level breakdowns across each EEG channel, and as you can see in the results for SynKI9-1(p60), the pipeline was producing clean, consistent HFO detections across all three EEG channels, which is exactly what we were hoping to see as the bug was fixed for running some of the longer EEG recordings of the mice.
After getting some of the detections running, I dove back into my literature review and read through a research paper that’s directly relevant to the experimental procedures we’re looking to apply for sleep-related activity detection and classification. The paper presents an EEG dataset recorded from nine mice going through a sleep deprivation paradigm, collecting nine days of continuous frontal and parietal EEG alongside accelerometer data, plus two hours of high-density EEG under both sleep-deprived and normal conditions. The researchers ran 18-hour sleep deprivation sessions across five consecutive days, and stored everything in a standardized BIDS format (Brain Imaging Data Structure) files like EDFs. This research also was quite important in helping me understand how to apply specific data analyses designed to help track oscillatory activity during sleep, things like slow waves and spindles, across different cortical regions and levels of sleep pressure, and they even included Python code for detecting those exact features. Across this literature review, I also learned about a three-channel EEG/EMG tethered mouse system that allows researchers to record three channels of EEG or EMG data simultaneously, which I realized directly connects to what we’ve been seeing in the PyHFO program outputs, where our recordings also come in across three EEG channels.
So this week’s blog covered how I was able to slice up some long EDF EEG segments using a short but effective shell script to get around a tricky processing bug, run the detectors manually, clean results at the bandpass filtering frequency, successfully save the detection outputs right alongside their source files in an organized way, and my main takeaways from my literature review. Then for next week, I’ll be diving into building a full EDF segmentation script that can handle creating many shorter and longer segments across a whole list of EDF files, and integrating Hydra for cleaner machine learning configuration management across our codebase. We’re getting much closer to having the full detection and classification workflow running end to end, and getting closer to finding significant patterns and potential biomarkers across the Syngap mice!

Leave a Reply
You must be logged in to post a comment.